1. Introduction
Protein structure prediction (PSP) methods developed in the field of computational biology aim to predict the three-dimensional structure of a protein from its amino acid sequence. This is essential for several applications, such as drug design and the design of novel enzymes1.
Profrager2 is a tool that supports PSP by implementing a strategy to reduce the conformational space complexity. This strategy consists of creating fragment libraries that can be used to construct segments of an amino acid target sequence. Profrager can be used to create customized protein fragment libraries with different sizes, and can be used in conjunction with other applications by several users.
The motivations to optimize Profrager include the following:
- It is a computationally intensive application.
- It can be used in conjunction with other applications; some scientific workflows developed and executed at LNCC (http://www.lncc.br/) are composed of tasks that execute Profrager.
- It can be used by several users across the Internet, because it is deployed in a web server.
Hybrid parallel computing architectures composed of multi-core and many-core processors based on Intel® Xeon® processors and Intel® Many Integrated Core Architecture have the potential to speed up computationally intensive applications in diverse fields. The contribution of this case study is to leverage knowledge about the efficient utilization of this hybrid parallel computing architecture by describing the optimization developed on the Profrager code.
The remainder of this white paper is organized as follows: Section 2 describes Profrager, section 3 shows the experiments setup, section 4 presents the performance analysis, section 5 describes the optimization developed, section 6 presents the evaluations, and section 7 presents the conclusion.
2. ProFrager
Profrager is implemented in C++ with Standard Template Library (STL) and with two main parts: input data reading and fragment library creation. The algorithms that compose each part are described in a high-level language:
- The first part (Algorithm 1) reads the following data that will be used during the execution:
- Input target sequence
- Secondary structure
- Structural geodatabase containing sequences and geometries from known proteins
- Structural database
- The second part (Algorithm 2 and Algorithm 3) is composed of a four-level nested loop. These loops create fragments and calculate two scores: score_matrix, based on structural database and score_psipred based on structural geodatabase and on the secondary structure. If the sum of these two scores is greater than the minimum score, the fragment is stored; otherwise, the fragment is discarded.
| 1) readDB(); 2) createGeoStructureList(); 3) read_FASTA_sequence(); 4) read_psipred(); |
Algorithm 1: Input data reading.
|
Algorithm 2: Main loop.
|
Algorithm 3: createFragments function.
The fragments of a library created by an execution of Profrager have a fixed size defined in the “fragment length” parameter. In general, applications that use libraries produced by Profrager need fragments of different sizes. In this sense, the experiments that use fragment libraries typically need to execute Profrager several times with different values for the “fragment length” parameter.
3. Setup of Experiments
To optimize Profrager we use an already-deployed hybrid parallel computing architecture composed of multi-core and many-core processors, and a workload with two examples.
The computing infrastructure is composed of a server deployed at NCC (http://www.unesp.br/portal#!/gridunesp) that has two Intel Xeon processors with hyper-threading enabled and four Intel® Xeon Phi™ coprocessors (Table 1).
|
|
Intel Xeon |
Intel Xeon Phi |
|---|---|---|
| Sockets x core x threads |
2 x 18 x 2 |
1 x 60 x 4 |
| Clock (Ghz) |
3 |
1 |
| L1 / L2 / L3 Cache (KB) |
32 / 256 / 46080 |
32 / 512 |
| DRAM (GB) |
128 |
16 |
Table 1: Description of computational infrastructure.
The workload is composed of two input data sequences developed by LNCC called T0793 and T0820, with 593 and 140 amino acids, respectively. The original version of Profrager is sequential and uses only one core for each execution. The results show that Profrager may have performance gains by fully utilizing all the computational resources provided by the server (Table 1).
| Data Set |
Execution Time (H:mm:ss) |
Standard Deviation (H:mm:ss) |
|---|---|---|
| T0793 |
0:28:28 |
0:01:04 |
| T0820 |
0:07:19 |
0:00:22 |
Table 2: Profrager execution time executing with the following input data sequence: T0793 and T0820 using the sequential version of Profrager.
4. Performance Analysis
Code optimization can be a challenging task due to the complexity of application code and due to the multi-level parallelism opportunities offered by hybrid parallel architectures. A performance analysis is essential to identify the optimization opportunities in the code and to provide information that can be used to guide the optimization. Typically, profilers are the tools that support this analysis, because they evaluate the code regions that are computationally intensive and also identify the possibilities to improve the performance of these regions3.
In order to execute the performance analysis of Profrager, we used Intel® Advisor, a profiler that supports developers in vectorizing code and developing parallelized code that uses threads. In this study case, we use two reports provided by Intel Advisor:
- Survey analysis: identifies the code regions that are computationally intensive and provides a complete vectorization report
- Suitability analysis: predicts the performance improvement that can be achieved by parallelizing loops
Sections 4.1, 4.2 and 4.3 show the steps performed in this analysis.
4.1 Project Creation
We created a new project in Intel Advisor using the following information: “application”, “application parameters”, and the source code folder (Figure 1).

Figure 1: Intel® Advisor project configuration.
4.2 Survey Analysis
We started the survey analysis after configuring the project (Figure 2).

Figure 2: Running the survey analysis.
The survey analysis shows the loops and functions in which an application spends the most time. This information is used to identify the opportunities for parallelization. We use the Total Time and Self Time:
- Total Time is the elapsed time in each loop considering the time involved in internal loops.
- Self Time is the elapsed time in each loop without internal loops.
The survey analysis of Profrager loops (Figure 3) indicates that loop 1 and loop 2 have Total Time close to the execution time of Profrager and no Self Time. That means that the body of these loops has a great amount of work performed by inner loops; therefore, it represents a good candidate for parallelization because it has high granularity.
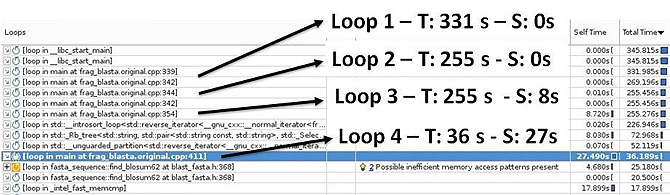
Figure 3: Self-time and Total time of functions and loops.
4.3 Suitability Analysis
The suitability analysis helps to quickly predict and to compare the parallel scalability and performance losses for different possible threading designs.
Parallel scalability prediction is based on annotations and a variety of modeling parameters. Annotations have to been inserted as pragmas in source code as follows:
- ANNOTATE_SITE_BEGIN(id): before beginning of loop
- ANNOTATE_ITERATION_TASK(id): first line inside the loop
- ANNOTATE_SITE_END(): after the end of loop
The loop in (Algorithm 4) illustrates how to insert annotations for suitability analysis.
| ANNOTATE_SITE_BEGIN( MySite1 ); for (int i = 0; i < j; i++) { ANNOTATE_ITERATION_TASK( MyTask1 ); c = A[i*n + j]; A[i*n + j] = A[j*n + i]; A[j*n + i] = c; } ANNOTATE_SITE_END(); |
Algorithm 4: Example of loop marked with Intel® Advisor annotations.
We added annotations into Profrager main loops, so that Intel Advisor can experiment with the potential parallelism of such loops. This helps determine the feasibility of adding parallelism, before you add parallel code (Figure 4).

Figure 4: Summary of annotations included in Profrager.
The Scalability estimate of loop 1 is shown in Figure 5. The Scalability estimate of loop 2 is shown in Figure 6. The Scalability results of loop 1 on the Intel Xeon Phi coprocessor are shown in Figure 7.
The analysis results provided by Intel Advisor indicates:
- Parallel version of loops 2, 3 and 4 present low performance gains (Figure 5). This result can be justified by those loops being executed several times and with a limited range of iterations. The time spent creating and destroying each thread is higher than executing the thread body.
- Loop 1 has the best scalability executing in the Intel Xeon processor (Figure 5) and the Intel Xeon Phi coprocessor (Figure 6).
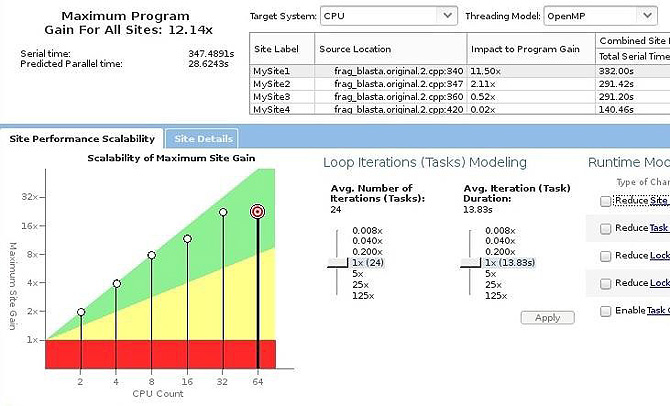
Figure 5: Scalability estimate of parallelizing loop 1 in the Intel® Xeon® processor.
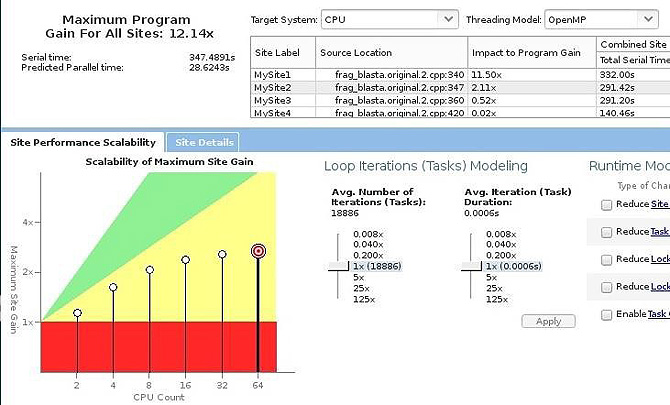
Figure 6: Scalability estimate of parallelizing loop 2 in the Intel® Xeon® processor.
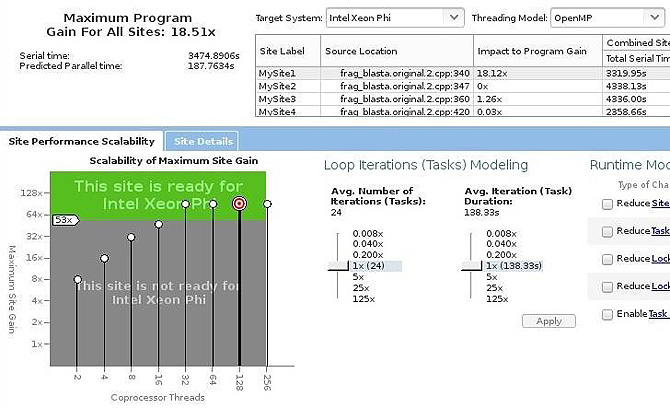
Figure 7: Scalability estimate of parallelizing loop 1 in the Intel® Xeon Phi™ coprocessor.
5. Optimization Development
We parallelized loop 1, based on the results of the suitability analysis, using the computational infrastructure and workload described in Section 4. In this sense, we developed two optimizations called Multi-threaded and OpenMP/MPI.
5.1 Multi-threaded
We developed a multi-threaded version of Profrager using OpenMP that runs in Intel Xeon processors. This optimization was conducted by including the OpenMP directive: “#pragma omp parallel for” directive on top of loop 1. In this context, the iterations of loop 1 are divided among threads provided by the system (Figure 8).
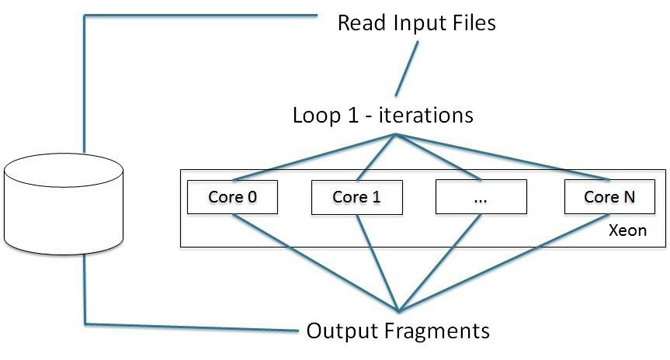
Figure 8: Multi-threaded for Intel® Xeon® processor optimization.Figure 8: Multi-threaded for Intel® Xeon® processor optimization.
5.2 OpenMP/MPI
We developed a parallel version of Profrager combining MPI and OpenMP to balance the load between processor and coprocessors. This strategy was implemented using the following structure (Figure 9):
- Start MPI rank 0 on Intel Xeon processor and one MPI rank at each Intel Xeon Phi coprocessor.
- Rank 0 performs Input Data Reading (Algorithm 1) and then sends all the data to other ranks using MPI_SEND.
- The execution of loop 1 steps (Algorithm 2 and Algorithm 3) are divided among ranks started at Intel Xeon Phi coprocessors.
- After finishing execution, the output files are sent to rank 0 on host via MPI_SEND. Rank 0 then starts to print the output results (line 6 of Algorithm 2)
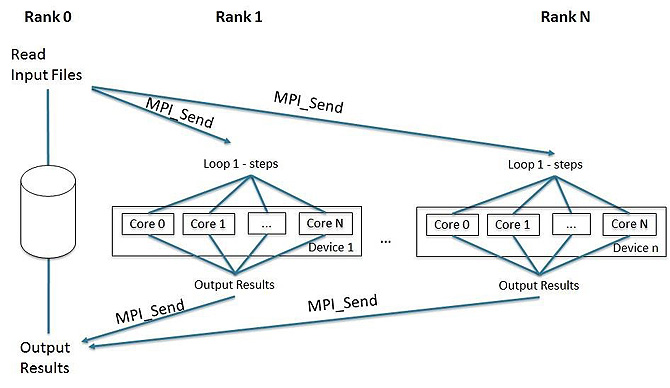
Figure 9: OpenMP/MPI for Intel® Xeon® processor and Intel® Xeon Phi™ coprocessor optimization.
6. Evaluation
In order to evaluate the optimizations developed and to compare against sequential version we performed the following tests:
- Execution of Multi-threaded version of one Profrager job varying the number of threads from 1 to 71 (Figure 10).
- Execution of OpenMP/MPI version of one Profrager job using one thread in Intel Xeon processor and several threads in Intel Xeon Phi coprocessors, varying the number of Intel Xeon Phi coprocessors from one to four (Figure 12).
- Execution of several Profrager jobs with different values for parameter “fragment length” (Figure 14) comparing the sequential version, Multi-threaded for Intel Xeon processor version and a load balancing strategy of executing half of the jobs using the Multi-threaded version and half of the jobs using the OpenMP/MPI version.
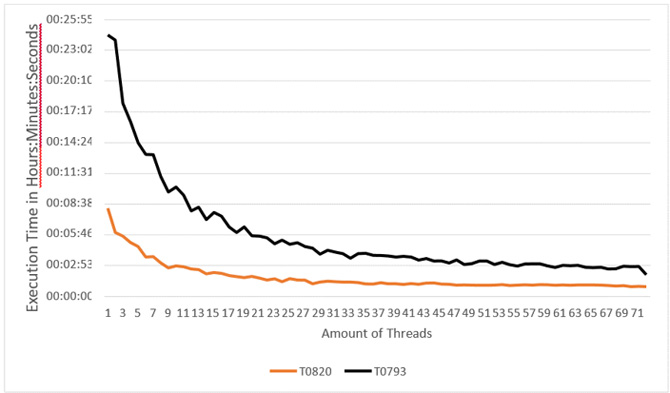
Figure 10: Evaluation of the multi-threaded Version of Profrager in Intel® Xeon® processor.
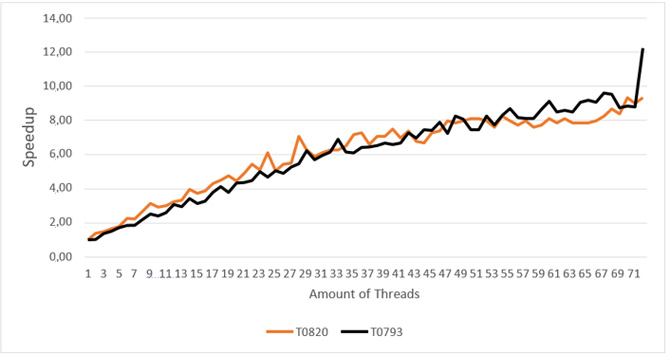
Figure 11: Speedup achieved by the multi-threaded version of Profrager in Intel® Xeon® processor.
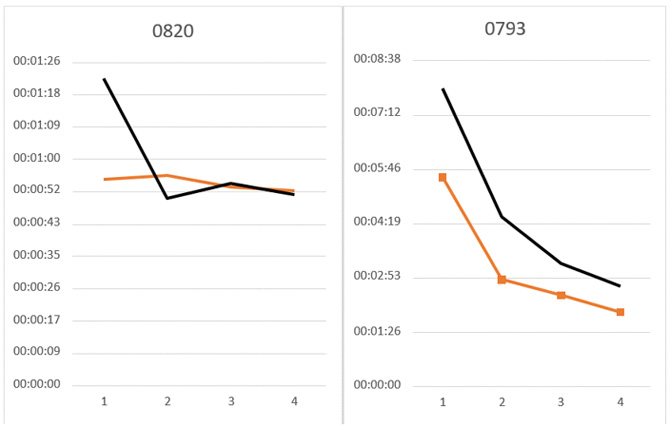
Figure 12: Evaluation of the OpenMP/MPI version in Intel® Xeon® processor and Intel® Xeon® Phi™ coprocessor.
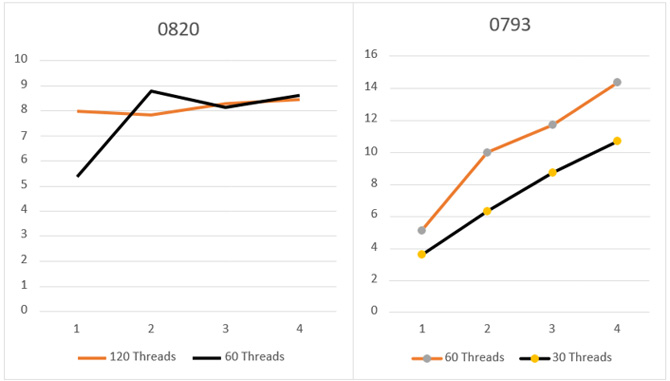
Figure 13: Speedup achieved by the OpenMP/MPI version in Intel® Xeon® processor and Intel® Xeon Phi™ coprocessor.

Figure 14: Several Profrager jobs execution in Intel® Xeon® processor and Intel® Xeon Phi™ coprocessor.

Figure 15: Speedup achieved executing several Profrager jobs in Intel® Xeon® processor and Intel® Xeon Phi™ coprocessor.
We will use speedup to compare the improvement of two optimizations against sequential versions because it indicates how much faster the optimized version is compared to the sequential version.
The tests executing one Profrager job using the Multi-threaded version showed high speedup for input target sequence T093 (12x) and T0820 (8x), using the maximum number of resources (Figure 11). Tests executing one Profrager job using the OpenMP/MPI version presented a similar speedup for input target sequence T093 (12x) and T0820 (8x) (Figure 13), using the maximum number of Intel Xeon Phi coprocessors.
The tests executing several Profrager jobs show lower speedup, because the concurrency for computational resources increased (Figure 15). Executing part of the jobs on the Intel Xeon processor using multi-threaded optimization and part of the jobs on Intel Xeon processor and Intel Xeon Phi coprocessor using OpenMP/MPI optimization showed better speedup, because this strategy enables the balancing of load using all the resources available on the node.
Other aspects observed during the optimization of Profrager are:
- The Intel Xeon Phi coprocessor presents memory constraints. In this case, Profrager experiments always had to be executed with less than 244 logical threads.
- I/O operations in Intel Xeon Phi coprocessors present low performance. In this case, we used MPI transfer instead of I/O operations.
- The MPI/OpenMP model can be used to port already parallelized applications to hybrid parallel architectures.
7. Conclusion
The optimization of Profrager described how to exploit the full capacity of hybrid parallel architectures with minimum modifications in the code.
This new version of Profrager presents the following advantages:
- Has the potential to increase the response time of diverse analysis executed by several users.
- Has only a few modifications in the code, which makes it easy to be managed by domain expert developers.
8. Future work
We have plans to work on two future works. The first one will parallelize other applications that compose LNCC workflows, and the second will explore the utilization of hybrid parallel architectures to minimize the execution time of workflows.
9. References
1 D. Baker and A. Sali, “Protein structure prediction and structural genomics.” Science, vol. 294, no. 5540, pp. 93–96. 2001.
2 Karina B. Santos, Raphael Trevizani, Fábio L. Custódio, and Laurent E. Dardenne, “Profrager Web Server: Fragment Libraries Generation for Protein Structure Prediction.” Biocomp. 2015.
3 Tomislav Janjusic, and Krishna Kavi, Chapter Three - Hardware and Application Profiling Tools, In: Ali Hurson, Editor(s), Advances in Computers, Elsevier, 2014, Volume 92, pages 105-160.
10. About Intel HPC
Supported by the Intel Modern Code Partner program, UNESP Center for Scientific Computing started in 2015 an action plan to establish a Parallel Programming Education Center focused on code modernization and dissemination of parallel processing concepts, aiming to collaborate on the transition to computing centered on parallelism. A series of training courses, workshops, and hands-on activities are being delivered targeting undergraduate and graduate students, academic researchers and practitioners, as well as engineers and developers from the industry sector.
Existing training material are being adapted, and newer ones being developed, to suit different needs according to the targeting audience, covering specialized areas of HPC and parallel computing with strong focus on parallelism and vectorization. The workshops cover a broad range of code modernization concepts and techniques, from the fundamentals of message-passing and shared-memory programming models through advanced performance optimization techniques for leading-edge HPC architectures.
Participants of the hands-on labs gain access to heterogeneous computing systems equipped with Intel Xeon processors and Intel Xeon Phi coprocessors, as well as Intel software development tools. Activities are centered in adapting and developing new course materials and providing practical training – with real-world examples and exercises – in parallel computing and related topics. Workshops are being delivered by experienced practitioners backed up by UNESP CSC engineering staff and UNESP / Intel Parallel Computing Center research fellows.
11. About The Authors
Silvio Stanzani (silvio@ncc.unesp.br) - Research Associate at NCC/UNESP
Raphael Cobe (rmcobe@ncc.unesp.br) - Research Associate at NCC/UNESP
Rogério Iope (rogerio@ncc.unesp.br) - Research Associate at NCC/UNESP
Igor Freitas (igor.freitas@intel.com) - HPC Application Engineer at Intel
Laurent Dardenne (dardenne@lncc.br) - Research Associate at the LNCC
Fábio Lima Custódio (flc@lncc.br) - Research Associate at the LNCC