Hello, fellow developers! I am Konstantinos, and I currently work at CERN as a research intern for this summer. I was accepted by the BioDynaMo (a.k.a Cells in the cloud) project, which is one of the projects that are part of the CERN openlab program.
For those who are not familiar with the CERN openlab program, it is a unique public-private partnership between CERN, research institutes, and pioneering ICT companies. Its primary goal is to accelerate the development of cutting-edge solutions which can influence the worldwide LHC community, as well as wider scientific research. Intel, among other contributors, is a leading supporter of this effort by providing its resources. If you are interested in learning more about the CERN openlab program, you can follow this link.
This year, since BioDynaMo is one of the projects sponsored by Intel that actively involve a summer intern, it takes part in The Modern Code Developer Challenge. You can check out the rest of the participating projects, by visiting the previous link. For that purpose, in the next few weeks, I will try to describe my work, the challenges I faced and share the overall experience, through a series of blog posts. In this very first post, I will give an overview of the project and elaborate on the current limitations.
Keep in mind that this is my first time blogging, so any comments are more than welcome! :)
Cells in the cloud
The BioDynaMo (Biological Dynamic Modeler) project aims at a general platform for computer simulations of biological tissue dynamics, such as brain development. It is a joint effort from between CERN, Newcastle University, Innopolis University, and Kazan Federal University. A possible (and desirable) use of the platform is the simulation of scenarios that can give researchers (e.g. neuroscientists) a better understanding of a wide variety of biological processes. Then, by using this new knowledge, it is possible to come up with better treatments.
If the above explanation seems somewhat complex, let's try something else; just take a look the following video:
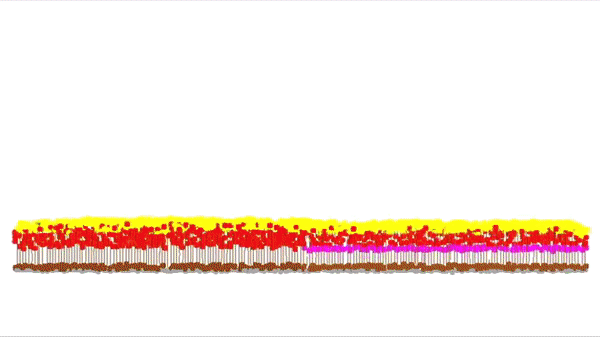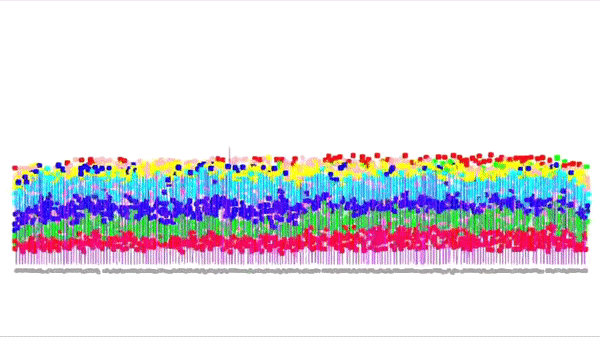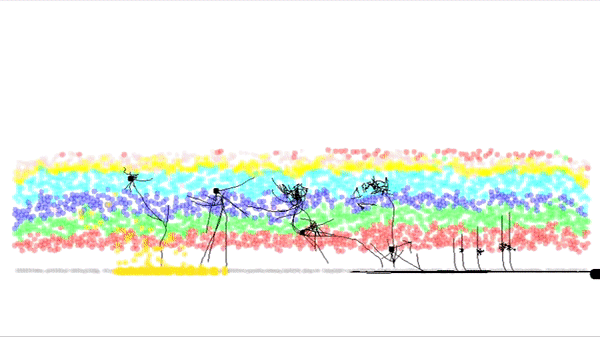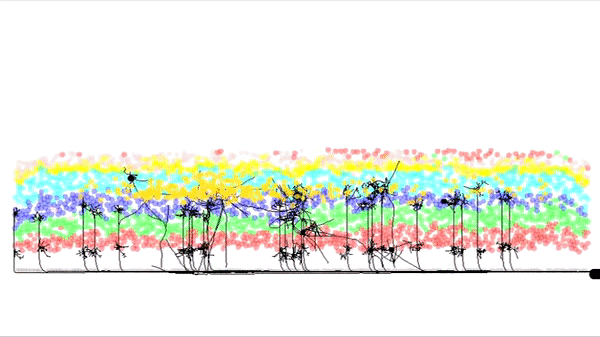
This is a simulation of a part of the cortex (specifically the outmost layer of the cortex). It was made by Andreas Hauri using Cx3D (you can find the video here). Behind this neat visualization, rather complicated computations are taking place, such as several physical and biological interaction forces. The BioDynaMo project focuses on the calculation of those forces, providing at the same time a unified, flexible, scalable and efficient platform for these kinds of (biological) simulations. Moreover, the platform intends to be accessible for researchers that don't have access to expensive HPC clusters.
Project history and limitations
The project started by porting algorithms written in Java* (originated from the Cx3D project), which didn't utilize recent technologies. During the last year, much work was carried out to implement a highly optimized multi-core version that can run efficiently on state-of-the-art systems, taking into account the processors architectures. The current implementation is written in modern C++ and uses OpenMP* framework for easy parallelization among the cores. Furthermore, the simulation state can be saved to disk, at any point during runtime, using a framework called ROOT (which is very popular among physicists). This prevents us from losing the state if some failure occurs in the system. Finally, there has been some work in progress for integrating with ParaView*, which is an open-source data analysis and powerful visualization software. (I promise to write a dedicated post for this!)
However, there is a major problem: Simulating large-scale complex biological systems require a tremendous amount of computing resources. For example, a simulation of just 10% of the cortex may require up to ~600TB of memory as well as calculating interactions among ~2 billion cells in every simulation step, depending on the complexity. It is quite obvious that these resources are not feasible on a single machine, as it is impossible to store this amount of data or process them efficiently.
- So, how can we tackle this problem?
You should have guessed the answer by now: we use distributed computing! Luckily for us, biological simulations, similar to physical ones, are parallelizable. So we can partition the computation into smaller parallel tasks and distributed them among our multiple nodes without much overhead. Then each node can independently execute its partial simulation and communicate with other nodes to retrieve the necessary data before the simulation can proceed to the next time step. In our case, we can apply the agent-based model since our simulations are only locally-coupled and in addition, the amount of data exchanged between nodes is bounded.
The goal for this summer!
Therefore, after pointing out the issue, my task is pretty straightforward: design and implement a distributed runtime prototype that can manage simple computations across many compute nodes. The runtime is intended to run both on high-performance clusters and cloud environments. Unfortunately, the two environments do not have many things in common. The former use tightly-coupled computers with very similar hardware, while the latter use loosely-coupled computers with heterogeneous hardware.
Running efficiently on both environments without much loss of performance and at the same time securing the reliable execution of the simulation is a very exciting and challenging problem! In the next blog post, I will describe in fine details the design process I followed for the distributed architecture and the compromises I had to make during that phase. Further, I will also introduce the tools, why I decided to use them and how they helped me overcome some fundamental issues.
Thanks for reading and stay tuned!