Introduction
This guide is targeted towards users who are already familiar with RELION* (REgularised LIkelihood OptimisatioN) and provides pointers and system settings for hardware and software that will provide the best performance for most situations. However, please note that we rely on the users to carefully consider these settings for their specific scenarios, since RELION can be deployed in multiple ways and this is a reference to one such use-case.
RELION is an open-source application suite used to discover the biological structures of molecules from data gathered via electron cryo-microscopy (cryo-EM). It is developed in the group of Sjors Scheres at the MRC Laboratory of Molecular Biology.
3rd Gen Intel® Xeon® Scalable processors deliver industry-leading, workload-optimized platforms with built-in AI acceleration, providing a seamless performance foundation to help speed data’s transformative impact, from the multi-cloud to the intelligent edge and back. Improvements of particular interest to RELION are:
- Enhanced Performance
- Increased DDR4 Memory Speed & Capacity
- Intel® Speed Select Technology
Tested hardware and software environment for this tuning guide:
We used the following hardware when writing this guide. It is not necessary to mirror our configuration, other than to get up-to-date Intel® Xeon® platforms with as many cores per node as you can afford.
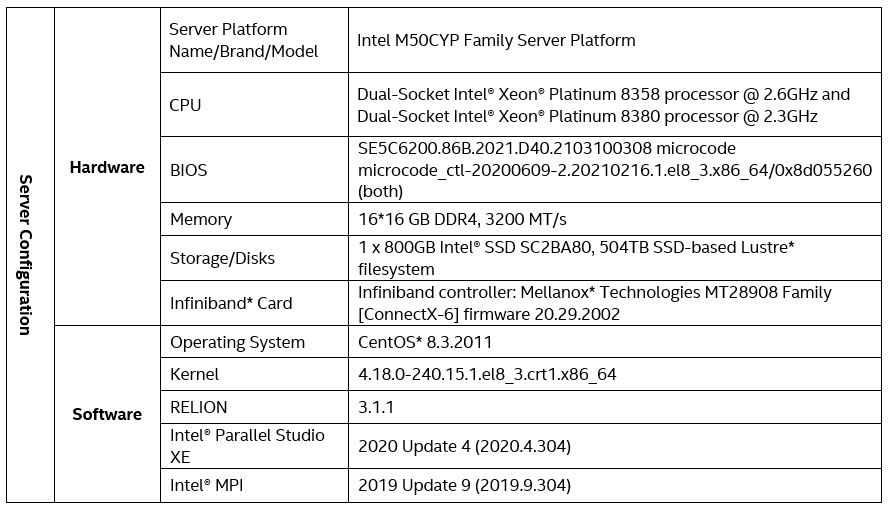
Hardware Tuning
BIOS Settings
For optimal RELION performance, make sure you have at least the following BIOS settings:
| Configuration Item | Recommended Value |
|---|---|
| Advanced/Power & Performance/CPU P State Control/CPU P State Control/Enhanced Intel SpeedStep(R) Tech | Enabled |
| Advanced/Power & Performance/CPU P State Control/CPU P State Control/Intel(R) Turbo Boost Technology | Enabled |
| Advanced/Processor Configuration/Intel(R) Hyper-Threading Tech | Enabled |
Description of Settings
Enhanced Intel SpeedStep® Technology
Enhanced Intel SpeedStep Technology allows the system to dynamically adjust processor voltage and core frequency, which can result in decreased average power consumption and decreased average heat production. In order that the Intel(R) Turbo Boost option to be available, Enhanced Intel SpeedStep Technology must be enabled.
Intel® Turbo Boost Technology
lntel Turbo Boost Technology allows the processor to automatically increase its frequency if it is running below power, temperature, and current specifications.
Intel® Hyper-Threading Technology
Intel Hyper-Threading Technology allows multithreaded software applications to execute two threads in parallel within each processor core. The result is that you can run threads on twice as many logical cores as physical cores in the system.
Memory Configuration/Settings
We recommend at least 256GB of memory per node. At least 1 DIMM per memory channel needs to be populated on your motherboard. Lower cpu-utilization could result if there are no DIMMs in a memory channel because of data contention. At this time, it does not appear that RELION workloads benefit from the use of persistent memory.
Storage/Disk Configuration/Settings
Datasets processed by RELION are large, and often benefit by processing on multiple nodes at once. As a result, we highly recommend using RELION on a cluster filesystem that is optimized for read/write accesses to both large and small files such as Lustre. Given the data requirements of CryoEM workloads, we recommend that your cluster filesystem be able to store at least 2PB per microscope.
Network Configuration/Setting
Best RELION performance is seen when the data sets are processed on multiple nodes using MPI. While RELION does not generate many MPI messages, RELION data sets and temporary files consume considerable disk space, and these files are accessed by all nodes during processing. RELION will *work* on a 100-base T network with a consumer network-attached filesystem, but we recommend an enterprise-class high-speed cluster fabric and parallel file system (the best you can reasonably afford). Cryo-EM microscopes are quite expensive, generate terabytes of output per day, and are often over-subscribed. You will need high-speed and reliable data handling to best access these scarce resources.
Software Tuning
Software configuration tuning is essential. From the Operating System to RELION’s configuration settings, they are all designed for general purpose applications and default settings are almost never tuned for best performance.
Linux Kernel Optimization Settings
<No specific workload setting for this topic>
RELION Execution Architecture
RELION consists of a suite of applications used to process Cryo-EM data in stages. Between each stage RELION reads the result of the last stage from disk and writes its results to disk at the end of a stage, often with intermediate results (to allow for resuming a stage from the middle). Stages may also be executed iteratively before proceeding to the next stage. A highly simplified view of this process flow is shown below:
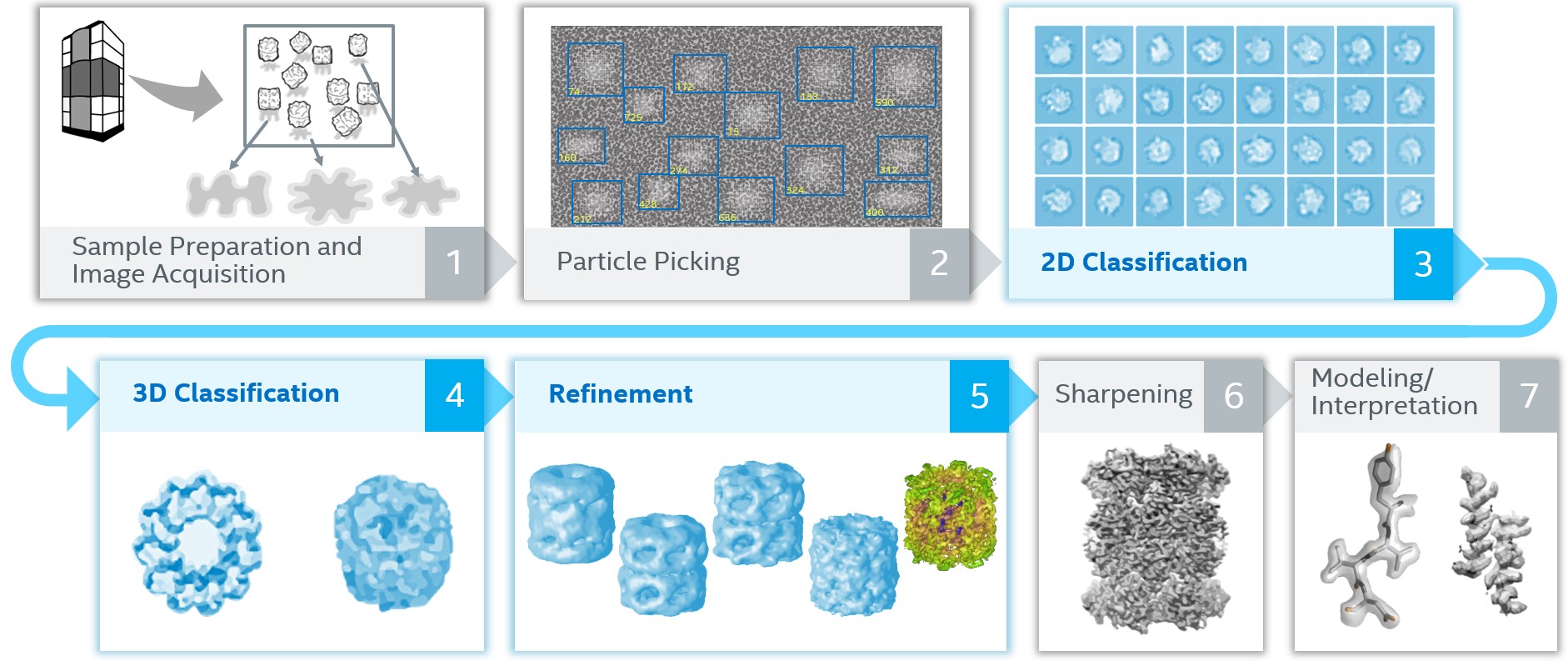
Figure 1. Simplified view of RELION process flow stages
The net effect of all these stages and intermediate files is that each Cryo-EM data set consists of (and generates) many large files. Rough processing times and storage requirements are shown here:
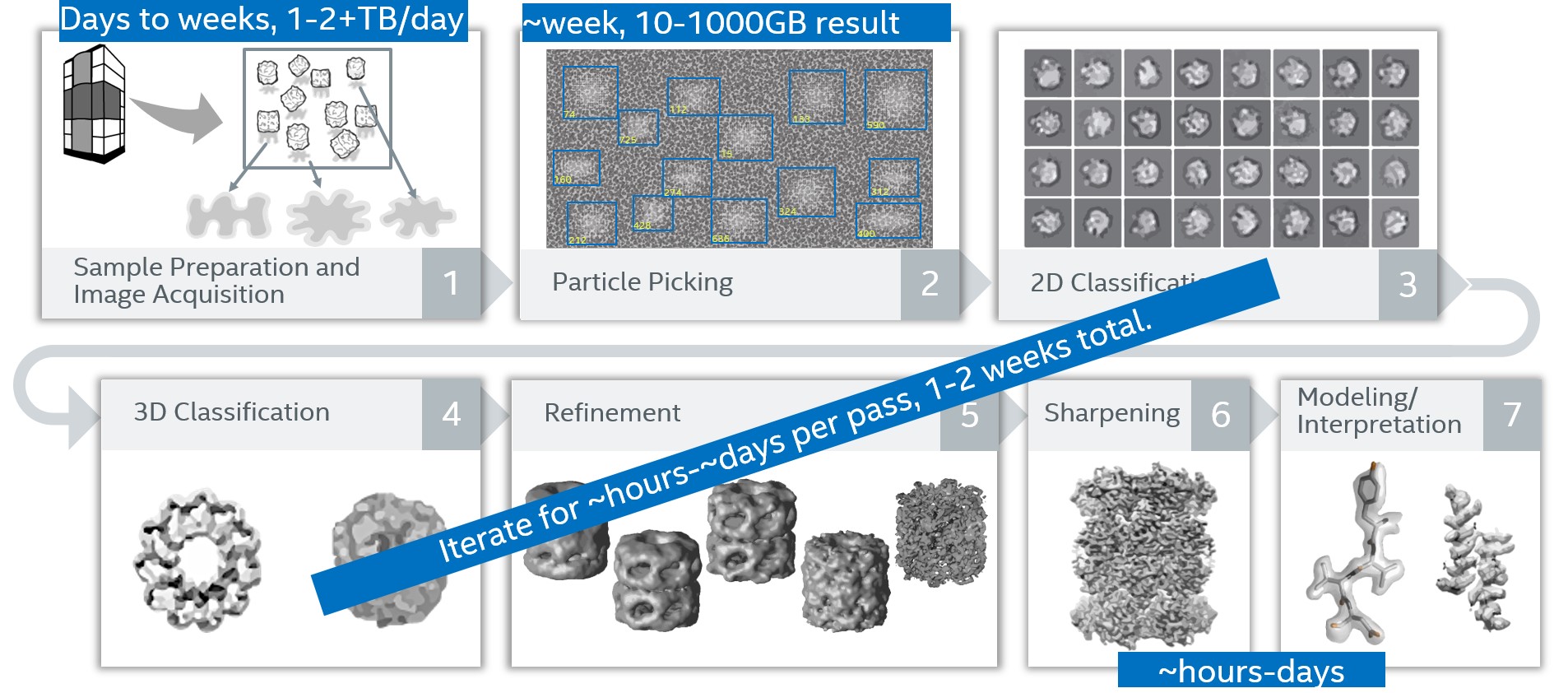

Figure 2. Simplified view of RELION process flow stages showing rough processing times and storage requirements
Once all the processing is complete, 100s of GB are typically archived for re-analysis, including sometimes the multi-terabytes of raw data.
These processing stages are highly compute and disk intensive. As a result, we recommend a minimum cluster for each microscope consisting of:
- 20 dual-socket compute nodes of 48-64 cores and 256-384GB (or more) memory per node
- A parallel file system of about 2PB of disk storage
- Nodes and parallel file system connected via a high-speed cluster fabric
These requirements scale linearly with each additional microscope.
Intel has worked with the creators of RELION to optimize stages 3-5 in the above figures. We will show you how to make use of these optimizations in the following section of this document.
RELION Install
The 2D classification, 3D classification, and 3D refinement algorithms in RELION have been optimized for Intel® Advanced Vector Extensions 512 (Intel® AVX-512) in the “Accelerated CPU" implementation, which is obtained by building RELION with the “ALTCPU=ON” CMAKE build flag, and then running with the “--cpu” option. These optimizations significantly improve performance on supported processors. They have been included in RELION since version 3.0, as built from its GitHub repository.
These optimizations may be disabled by running without the --cpu option, or by running with the --gpu option. In all cases, better performance has been observed when using the free Intel® C++ Compiler over the default Linux* compiler.
In general, the build and run procedure is unchanged from typical RELION procedures beyond the above-mentioned build and runtime flags. For clarify, however, here is the build process:
1) Download RELION (with the “3.1.1” or later build tag)
2) Setup the compiler and library environment. Here, we use the Intel® MPI Library, Intel® Math Kernel Library (Intel® MKL), and the Intel® C++ Compiler. For Intel® Parallel Studio XE, the environment can be set up as follows. The example below is for Intel® Parallel Studio XE 2020:
$ source /opt/intel/impi/2019.9.304/intel64/bin/mpivars.sh intel64
$ source /opt/intel/compilers_and_libraries_2020.4.304/linux/bin/compilervars.sh intel64
$ source /opt/intel/compilers_and_libraries_2020.4.304/linux/mkl/bin/mklvars.sh intel64
3) Build a version of RELION for Intel® Xeon® Scalable Processors with AVX-512 from the “relion” directory that results when you pull the source from git:
$ mkdir build
$ cd build
$ cmake -DCMAKE_C_COMPILER=icc -DCMAKE_CXX_COMPILER=icpc -DMPI_C_COMPILER=mpiicc -DMPI_CXX_COMPILER=mpiicpc -D CMAKE_C_FLAGS="-O3 -ip -g -debug inline-debug-info -xCOMMON-AVX512 -qopt-report=5 -restrict " -D CMAKE_CXX_FLAGS="-O3 -ip -g -debug inline-debug-info -xCOMMON-AVX512 -qopt-report=5 -restrict " -DALTCPU=ON -DMKLFFT=ON -DGUI=OFF -D CMAKE_BUILD_TYPE=Release ..
$ make -j 20
$ cp bin/relion_refine_mpi ../relion_refine_mpi_3.1.1_AVX512
4) (Optional) build a version of RELION for non-Intel processors with AVX-2 from the “relion” directory that results when you pull the source from git:
$ cd ..
$ rm -rf build
$ mkdir build
$ cd build
$ cmake -DCMAKE_C_COMPILER=icc -DCMAKE_CXX_COMPILER=icpc -DMPI_C_COMPILER=mpiicc -DMPI_CXX_COMPILER=mpiicpc -D CMAKE_C_FLAGS="-O3 -ip -g -debug inline-debug-info -march=core-avx2 -qopt-report=5 -restrict " -D CMAKE_CXX_FLAGS="-O3 -ip -g -debug inline-debug-info -march=core-avx2 -qopt-report=5 -restrict " -DGUI=OFF -DALTCPU=ON -DMKLFFT=ON -D CMAKE_BUILD_TYPE=Release ..
$ make -j 20
$ cp bin/relion_refine_mpi ../relion_refine_mpi_3.1.1_nonIntelAVX2
To verify that we have successfully built and run RELION, we use the Plasmodium Ribosome data set, and run it with only minor changes to the documented processing parameters (more on that below).
Generic instructions for downloading the workload and running it can be found on the Relion benchmark & computer hardware webpage. Note that you will need 47 gigabytes for the initial benchmark data, and a similar amount for the results generated while running the benchmark. We recommend the data be put on a fast cluster file system.
RELION Tuning
Best performance is seen when running RELION with Intel® Hyper-Threading Technology enabled, and 8 MPI compute ranks per node plus one (for the data sets tested to date). The “plus one” comes from the fact that the coordinating rank uses as many threads as all the other ranks, but does no processing while they are running, so we can have a second rank use the same logical cores as one compute rank.
We don’t recommend using more than 16 machines total for a given RELION job at this time, as the scaling appears limited. This of course depends on the size of the input data, but RELION's data-flow management is not equipped to handle arbitrarily large jobs, so scaling will likely be limited both by available task parallelism AND latency at this point.
Where RELION parameters are concerned, we recommend:
- Since we see best results with 8 MPI compute ranks per node on dual-socket systems with Intel® processors, --j should be set to the total logical cores (with Intel Hyper-Threading Technology enabled) in the machine divided by the number of compute ranks per machine (8)
- We recommend running with more than 9 threads per rank (--j) if at all possible, which may mean dropping to a smaller number of ranks per node.
- Clusters with less than 128GB of RAM per node will want to run with fewer ranks per system to minimize cases where they run out of system memory or swap.
- Set pool size (--pool) to roughly twice the number of threads (--j), and not lower than about 30 (40 seems to work well). However, lowering the pool size may also decrease the memory used by a process or rank if that is an issue.
Systems with 256GB or more are recommended for the CPU-accelerated kernels.
We recommend setting the following environment variables:
$ export OMP_SCHEDULE="dynamic"
$ export KMP_BLOCKTIME=0
$ export I_MPI_PIN_DOMAIN=<the total logical cores in the machine divided by the number of compute ranks per machine>
$ export I_MPI_DEBUG=5
$ export I_MPI_FABRICS=<appropriate fabrics for your installation>
Single-node example
If we run RELION on a dual-socket system consisting of 8358 Intel® Xeon® Scalable Processors with Intel Hyper-Threading Technology enabled, we have a total of 128 logical cores. At 8 compute ranks per node, that gives us 16 threads per rank, running with a pool size of 40. And an additional rank for the controlling rank. The resulting command sequence would look something like this:
$ export OMP_SCHEDULE="dynamic"
$ export KMP_BLOCKTIME=0
$ export I_MPI_PIN_DOMAIN=16
$ export I_MPI_DEBUG=5
$ export I_MPI_FABRICS=<appropriate fabrics for your installation>
$ mpirun -n 9 relion_refine_mpi <other flags and options> --j 16 --pool 40
Multi-node example
If we run RELION on four dual-socket systems consisting of 8380 Intel® Xeon® Scalable Processors with Intel® Hyper-Threading Technology enabled, we have a total of 160 logical cores per node. At 8 compute ranks per node, that gives us 20 threads per rank, running with a pool size of 40. There will be a total of 33 ranks since we need to add one for the coordinating rank to the total. The resulting command sequence would look something like this:
$ export OMP_SCHEDULE="dynamic"
$ export KMP_BLOCKTIME=0
$ export I_MPI_PIN_DOMAIN=20
$ export I_MPI_DEBUG=5
$ export I_MPI_FABRICS=<appropriate fabrics for your installation>
$ mpirun -n 33 relion_refine_mpi <other flags and options> --j 20 --pool 40
Best Practices for Testing and Verification
To make sure RELION is built optimally for 3rd Gen Intel® Xeon® Scalable processors, and to identify any cluster or machine issues that might affect cluster performance, we like to use the Plasmodium Ribosome workload described in the introduction to benchmark the result.
As noted before, best performance is seen when running RELION with Intel Hyper-Threading Technology enabled, so please set your benchmark system(s) up that way.
After downloading and unpacking the Plasmodium Ribosome data set as instructed below, go to the top level of the resulting directory tree. You should find a file called emd_2660.map and a directory called Particles. We will show you how to run the data set from this location.
One-time setup
The Plasmodium Ribosome data set has been around for some time, and RELION has evolved during that time. Before you run the benchmark, you should convert the data file listing the particles to process from an old to new format (RELION can still read the old format – it will just complain and do unnecessary extra work):
$ <relion_source_tree/build/bin/>relion_convert_star --i Particles/shiny_2sets.star --o
Particles/shiny_2sets_3.1.star
With this out of the way, we can run the benchmark
Timing your benchmarks
RELION has built-in timer code you can enable by building using the “Benchmarking” rather than “Release” for CMAKE_BUILD_TYPE, but to prevent it from interfering with the overall runtime or scalability of the application, we recommend not using it. Instead, you can try to time the binary with /usr/bin/time. A more friendly technique that produces times in seconds that are easy to add to spreadsheets, is to bracket the mpirun command as follows in a script:
btime=`date +%s`
<run_RELION_as_per_below_via_mpirun>
etime=`date +%s`
diff=`expr $etime - $btime`
echo "RELION ... < information about the run parameters> completed in the following time (seconds): $diff "
Testing on dual-socket systems consisting of 8358 Intel® Xeon® Scalable Processors with Intel® Hyper-Threading Technology enabled
As you may recall from our above example, a single node of such a dual-socket system has a total of 128 logical cores. At 8 compute ranks per node, that gives us 16 threads per rank, running with a pool size of 40.
To run the Plasmodium Ribosome data set on a single node under Intel IMPI with 8 compute rank plus one coordinating rank, you would issue the following commands from the top level of the unpacked data set:
$ mkdir Results
$ export OMP_SCHEDULE="dynamic"
$ export KMP_BLOCKTIME=0
$ export I_MPI_PIN_DOMAIN=16
$ export I_MPI_FABRICS=<appropriate fabrics for your installation>
$ btime=`date +%s`
$ mpirun -hostfile <one_node_host_file> -env OMP_SCHEDULE=dynamic -env KMP_BLOCKTIME=0 -perhost 9 -np 9 <binary_location>/relion_refine_mpi_3.1.1_AVX512 --i Particles/shiny_2sets_3.1.star --ref emd_2660.map:mrc --firstiter_cc --ini_high 60 --dont_combine_weights_via_disc --ctf --ctf_corrected_ref --tau2_fudge 4 --particle_diameter 360 --K 6 --flatten_solvent --zero_mask --oversampling 1 --healpix_order 2 --offset_range 5 --offset_step 2 --sym C1 --norm --scale --random_seed 0 --pool 40 --j 16 --iter 25 --cpu --o Results/cpu3d
$ etime=`date +%s`
$ diff=`expr $etime - $btime`
echo "RELION 3.1.1 completed 25 iterations of Plasmodium Ribosome using 9 ranks and 16 threads per rank and a pool size of 40 on a single Intel® Xeon® 8358 Scalable Processor node in the following time (seconds): $diff "
To run this data set on four nodes of such dual-socket systems, we would again have eight compute ranks per node for a total of 32, plus one rank for coordination. The resulting benchmark run would look as follows:
$ mkdir Results
$ export OMP_SCHEDULE="dynamic"
$ export KMP_BLOCKTIME=0
$ export I_MPI_PIN_DOMAIN=16
$ export I_MPI_DEBUG=5
$ export I_MPI_FABRICS=<appropriate fabrics for your installation>
$ btime=`date +%s`
$ mpirun -hostfile <four_node_host_file> -env OMP_SCHEDULE=dynamic -env KMP_BLOCKTIME=0 -perhost 8 -np 33 <binay_location>/relion_refine_mpi_3.1.1_AVX512 --i Particles/shiny_2sets_3.1.star --ref emd_2660.map:mrc --firstiter_cc --ini_high 60 --dont_combine_weights_via_disc --ctf --ctf_corrected_ref --tau2_fudge 4 --particle_diameter 360 --K 6 --flatten_solvent --zero_mask --oversampling 1 --healpix_order 2 --offset_range 5 --offset_step 2 --sym C1 --norm --scale --random_seed 0 --pool 40 --j 16 --iter 25 --cpu --o Results/cpu3d
$ etime=`date +%s`
$ diff=`expr $etime - $btime`
echo "RELION 3.1.1 completed 25 iterations of Plasmodium Ribosome using 33 ranks and 16 threads per rank and a pool size of 40 on four Intel® Xeon® 8358 Scalable Processor nodes in the following time (seconds): $diff "
Testing on dual-socket systems consisting of 8380 Intel® Xeon® Scalable Processors with Intel® Hyper-Threading Technology enabled
As you may recall from our above example, a single node of such a dual-socket system has a total of 160 logical cores. At 8 compute ranks per node, that gives us 20 threads per rank, running with a pool size of 40.
We’d run the Plasmodium Ribosome data set as above, except that we would set I_MPI_PIN_DOMAIN to 20 rather than 16, and similarly change the “--j” parameter (which represents the number of threads for RELION to use per rank) to 20 rather than 16.
Comparing to the “Reference” results
If you want to compare the results of a RELION run with the “--cpu” or “--gpu” options to the original, high-precision (but slow) RELION algorithms using a tool like Chimera, it’s simple. Just run as above without the “--cpu” option in a different output directory or with a different output name. The resulting *.mrc files for the run with the “--cpu” or “--gpu” option and the run without either option (the “reference results”) can loaded in Chimera and overlaid to give you an idea of how the results differ. You can also run 3D refinement and compare the Fourier Shell Correlation curves.
Conclusion
RELION can process a wide range for data sets in many ways on multiple hardware architectures. The above guidelines should give you a good idea for how to run RELION on 3rd Gen Intel® Xeon® Scalable processor-based systems to get the best performance. They also outline a general-purpose cluster architecture that should excel when processing other classes of workloads, as well as Cryo-EM . With the 3rd Generation Intel® Xeon® Scalable processor, Intel takes it even further by optimizing the platform as a whole -- CPU, memory, storage, and networking working together for the best user experience.
Related Tools
RELION includes a suite of tools that can be used to set up and run the various RELION stages, verify its results, etc. See the RELION documentation (in particular the tutorial) for details. In addition to those changes, we recommend downloading the Chimera tool from the Resource for Biocomputing, Visualization, and Informatics group at the University of California, San Francisco. This tool can be used to visualize and compare the results of RELION runs.
Additional Resources
- Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2
- Cryo-EM structure of the activated NAIP2-NLRC4 inflammasome reveals nucleated polymerization
- Structural mechanism for nucleotide-driven remodeling of the AAA-ATPase unfoldase in the activated human 26S proteasome
References
-
For more information, see “BIOS Setup Utility User Guide for the Intel® Server Board D50TNP and M50CYP Family Supporting the 3rd Generation Intel® Xeon® Scalable Processor Family”
Feedback
We value your feedback. If you have comments (positive or negative) on this guide or are seeking something that is not part of this guide, please reach out and let us know what you think.
Notices & Disclaimers
Intel technologies may require enabled hardware, software or service activation.
No product or component can be absolutely secure.
Your costs and results may vary.
Code names are used by Intel to identify products, technologies, or services that are in development and not publicly available. These are not "commercial" names and not intended to function as trademarks
The products described may contain design defects or errors known as errata which may cause the product to deviate from published specifications. Current characterized errata are available on request.
© Intel Corporation. Intel, the Intel logo, and other Intel marks are trademarks of Intel Corporation or its subsidiaries. Other names and brands may be claimed as the property of others.